Cellular Barcoding Used to Characterize Clones of Breast Tumor Cells
By LabMedica International staff writers
Posted on 12 Mar 2019
Cancer researchers used an advanced cellular barcoding technique to characterize cells in primary and disseminated tumors in mouse models of human triple-negative breast cancer.Posted on 12 Mar 2019
Triple-negative breast cancer (TNBC) refers to any breast cancer that does not express the genes for estrogen receptor (ER), progesterone receptor (PR), and HER2/neu. Lack of these receptors increases the difficulty of treating the disease, since most hormone therapies target one of the three receptors. Primary triple negative breast cancers are prone to dissemination but sub-clonal relationships between tumors and resulting metastases are poorly understood.
Image: Cellular barcoding may help to better understand how breast cancer evolves and how treatment strategies can be refined for increased benefits (Photo courtesy of Breast Cancer News).
To help clarify this relationship, investigators at the Walter and Eliza Hall Institute of Medical Research (Melbourne, Australia) worked with mouse models carrying two treatment-naïve TNBC patient-derived xenografts (PDXs) in order to track the fate of thousands of barcoded clones in primary tumors, and their metastases.
The investigators sought to capture the majority of tumor biomass and tumor cells at distal sites, and examine their heterogeneity in an unbiased fashion. Distal sites included cells that had shed from the primary tumor and were found in the blood stream as circulating tumor cells (CTCs), or in other distal organs (e.g. lung, bone marrow, ovaries, and kidney) where they accumulated as disseminated tumor cells (DTCs).
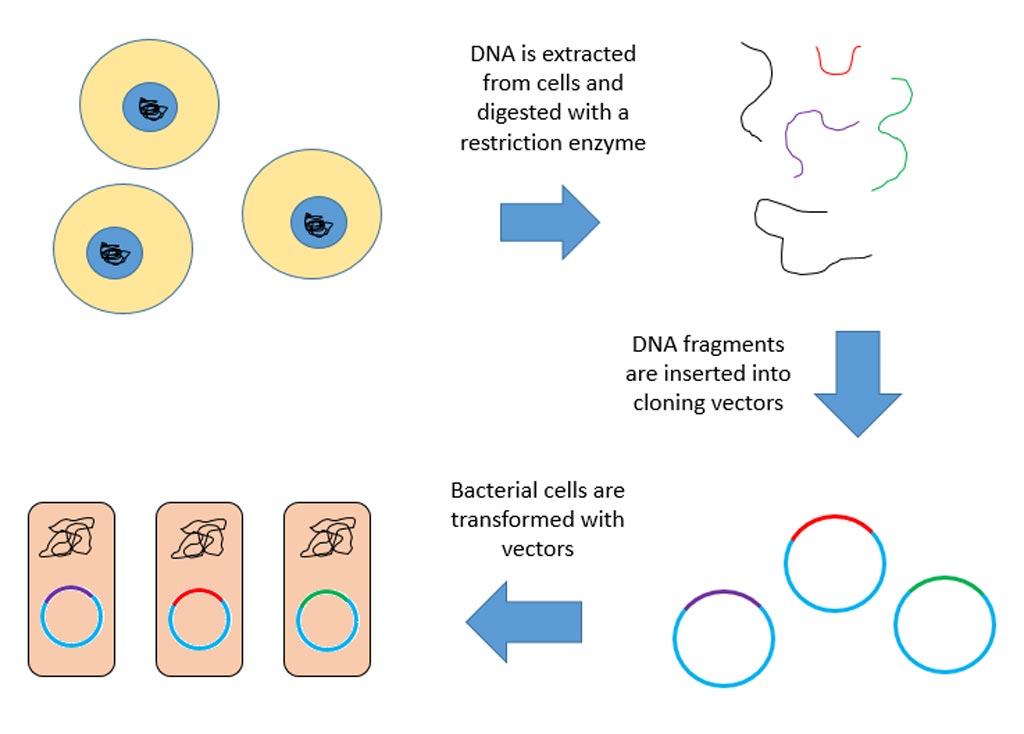
To accomplish this task, the investigators utilized cellular barcoding, which allowed robust assessment of clonal diversity and numbers at high resolution and depth, including confident detection of clones as small as five to 10 cells amongst millions.
The investigators reported in the February 15, 2019, online edition of the journal Nature Communications that tumor resection had a major impact on reducing clonal diversity in secondary sites, indicating that most disseminated tumor cells lacked the capacity to "seed", and therefore originated from "shedders" that did not persist. The few clones that continued to grow after resection ("seeders") did not correlate in frequency with their parental clones in primary tumors. Cisplatin treatment of one BRCA1-mutated PDX model had a surprisingly minor impact on clonal diversity in the relapsed tumor yet purged 50% of distal clones. Therefore, clonal features of shedding, seeding, and drug resistance are important factors to consider for the design of therapeutic strategies.
"The barcoding technique," said first author Dr. Delphine Merino, head of the tumor progression and heterogeneity laboratory, at the Olivia Newton-John Cancer Research Institute (Melbourne, Australia), "enabled us to identify the clones that were able to get into the bloodstream and make their way into other organs where they would "seed" new tumor growth. Our study revealed that only a select few clones were actually responsible for the metastasis."
Related Links:
Walter and Eliza Hall Institute of Medical Research
Olivia Newton-John Cancer Research Institute