Mutations Force Proteins to Adopt Solid-like Structures
By LabMedica International staff writers
Posted on 31 Jan 2018
A recently published paper provided a unified structural view of self-assembly, aggregation, and interactions of the hnRNPA2 protein and the distinct effects of small chemical changes such as disease mutations and arginine methylation on these assemblies.Posted on 31 Jan 2018
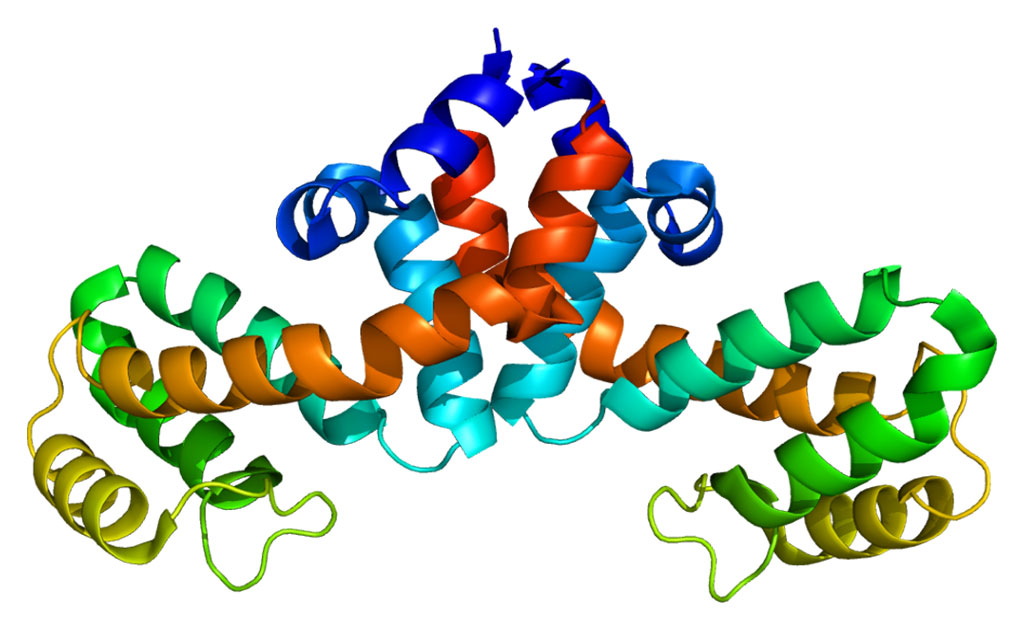
The hnRNPA2 protein is a component of RNA-processing organelles that lack membranes. This protein forms inclusions when mutated in a syndrome characterized by the degeneration of neurons (bearing features of amyotrophic lateral sclerosis [ALS] and frontotemporal dementia), muscle, and bone.
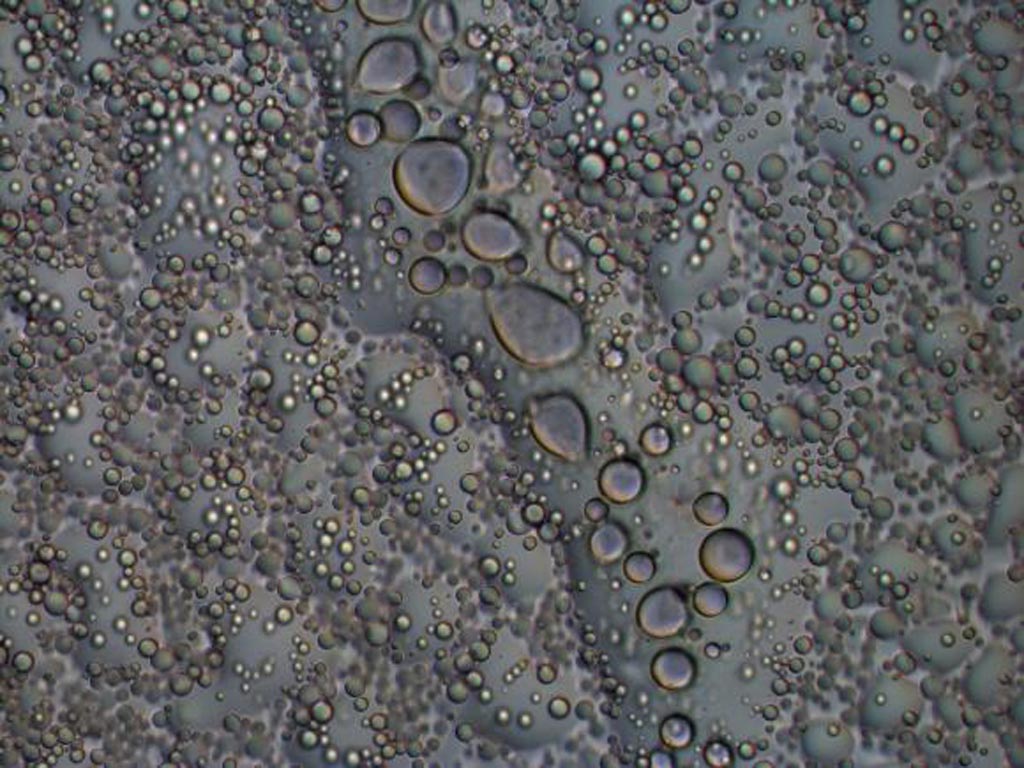

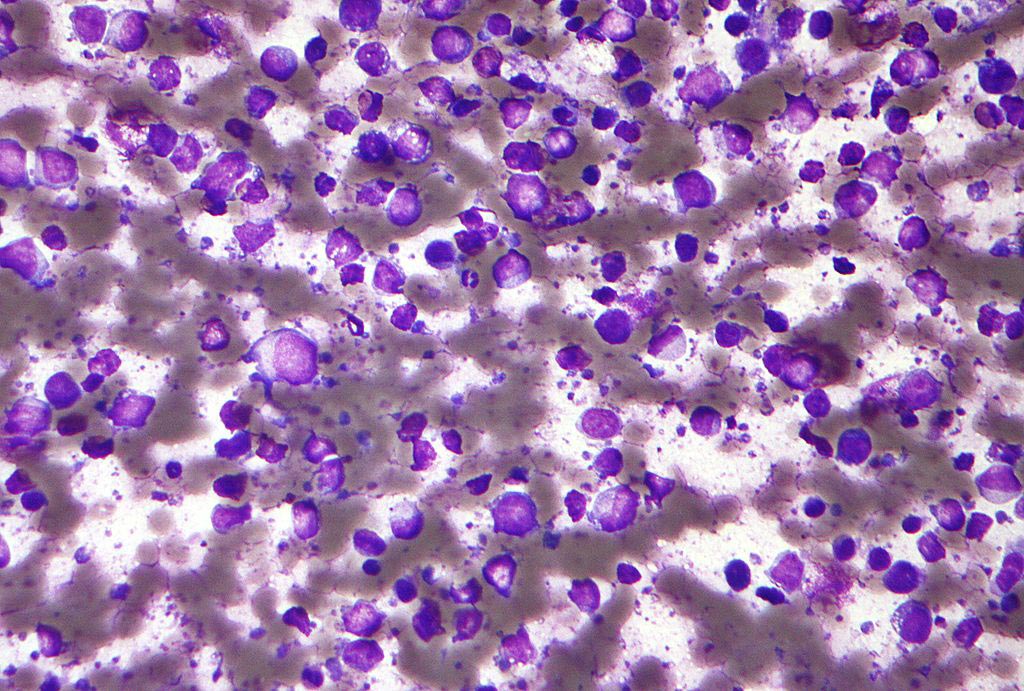
Image: The hnRNPA2 protein forms liquid droplets in a test tube as seen by light microscopy. These structures allow researchers to examine how disease mutations and functional modifications change the behavior of the proteins with atomistic detail (Photo courtesy of Veronica Ryan, Brown University).
In the January 18, 2018, online edition of the journal Molecular Cell investigators at Brown University (Providence, RI, USA) used nuclear magnetic resonance (NMR) spectroscopy, computer simulations, and microscopy to show how mutations and arginine methylation, a functional modification common to a large family of proteins with low-complexity domains, altered the formation low-complexity protein liquid droplets and their conversion to solid-like states in disease situations.
"We show how small chemical changes - involving only a few atoms - lead to big changes in assembly and disease-associated aggregation," said senior author Dr. Nicolas Fawzi, assistant professor of molecular pharmacology, physiology, and biotechnology at Brown University.
"These interactions are more dynamic and less specific than previously thought. A molecule does not take just one shape and bind to one shape but a molecule is flexible and interacts in flexible ways. Because these low-complexity domains are too flexible to be directly targeted by standard drugs, finding out how cells use and tame these domains is a potential route to stopping their unwanted assembly in disease."
Related Links:
Brown University