Advanced Proteomics Strategy Identifies Promising Drug Target
By LabMedica International staff writers
Posted on 10 Oct 2017
Use of an advanced proteomics strategy led to the discovery of a protein in non-small-cell lung cancer (NSCLC) cells that could be targeted by low molecular weight drugs.Posted on 10 Oct 2017
The transcription factor NRF2 (Nuclear factor (erythroid-derived 2)-like 2) is a master regulator of the cellular antioxidant response, and it is often genetically activated in NSCLCs by, for instance, mutations in the negative regulator KEAP1 (Kelch-like ECH-associated protein 1). While direct drug-based inhibition of NRF2 has not been possible (NRF2 regulates the activities of genes expressed in cell types throughout the body, so a powerful NRF2-blocking agent would have excessive side effects), its overexpression modifies biochemical networks in cancer cells in a fashion that may create special vulnerabilities.
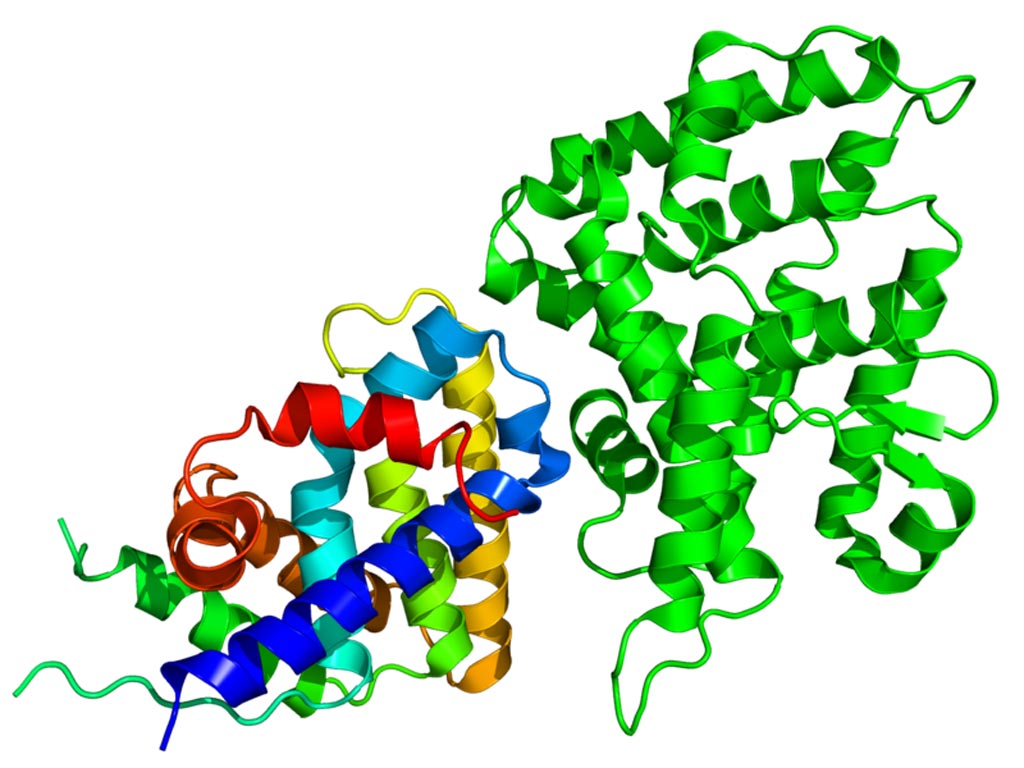

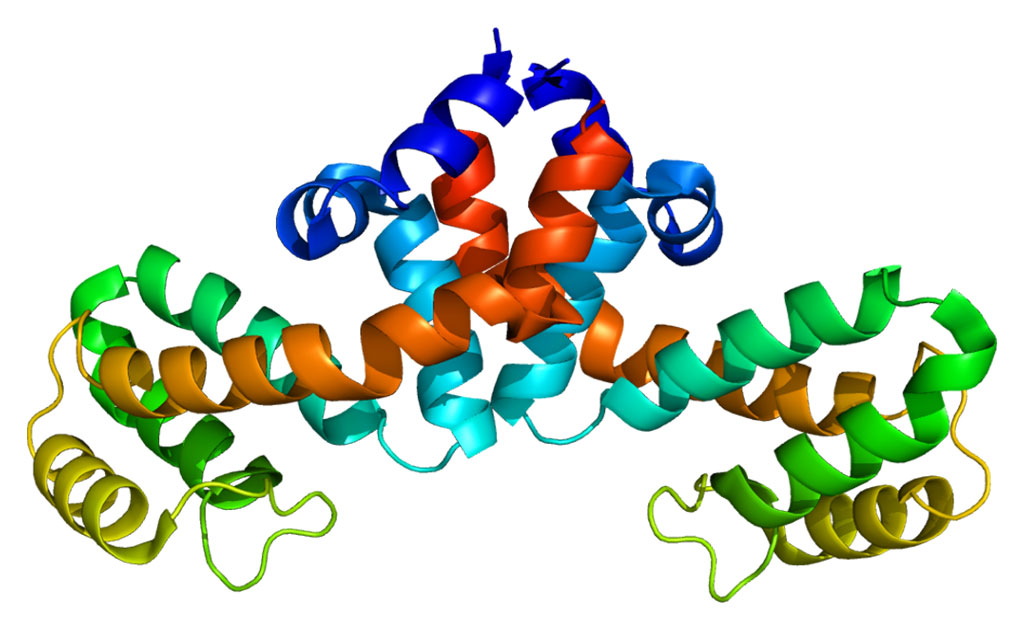
Image: The Crystallographic structure of NR0B1 (rainbow colored) complexed with the nuclear receptor protein LRH-1 (Liver receptor homolog-1) (Photo courtesy of Wikimedia Commons).
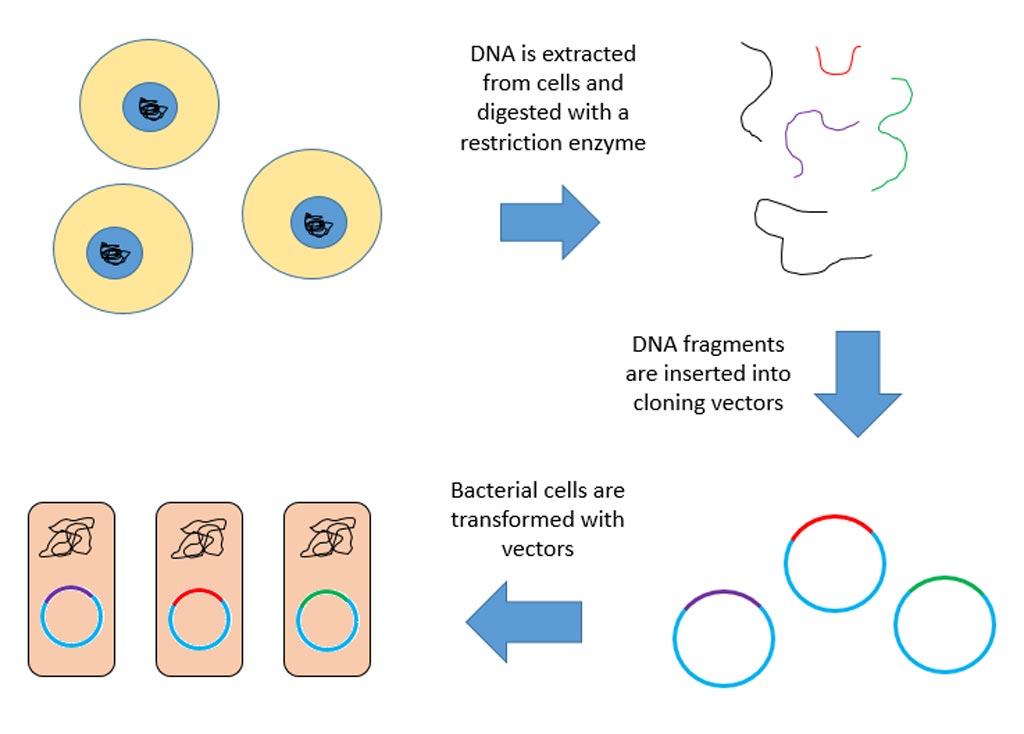
Investigators at The Scripps Research Institute (La Jolla, CA, USA) used chemical proteomics to map druggable proteins that were selectively expressed in KEAP1-mutant NSCLC cells. They reported in the September 28, 2017, online edition of the journal Cell that principal among these proteins was NR0B1, an atypical orphan nuclear receptor that was shown to engage in a multimeric protein complex to regulate the transcriptional output of KEAP1-mutant NSCLC cells. NR0B1 is located normally in the nucleus in lung cancer cells as a component of a larger protein complex that regulates gene expression.
The investigators identified small molecules that covalently targeted a conserved cysteine within the NR0B1 protein interaction domain, and they demonstrated that these compounds disrupted NR0B1 complexes and impaired the anchorage-independent growth of KEAP1-mutant cancer cells.
"This new approach shows promise for identifying previously unrecognized druggable targets in cancers that lack effective treatments," said senior author Dr. Benjamin F. Cravatt, professor of molecular medicine at The Scripps Research Institute.
Related Links:
The Scripps Research Institute