X-Ray Crystallography Findings Aid Discovery of New Drugs
By LabMedica International staff writers
Posted on 18 Apr 2017
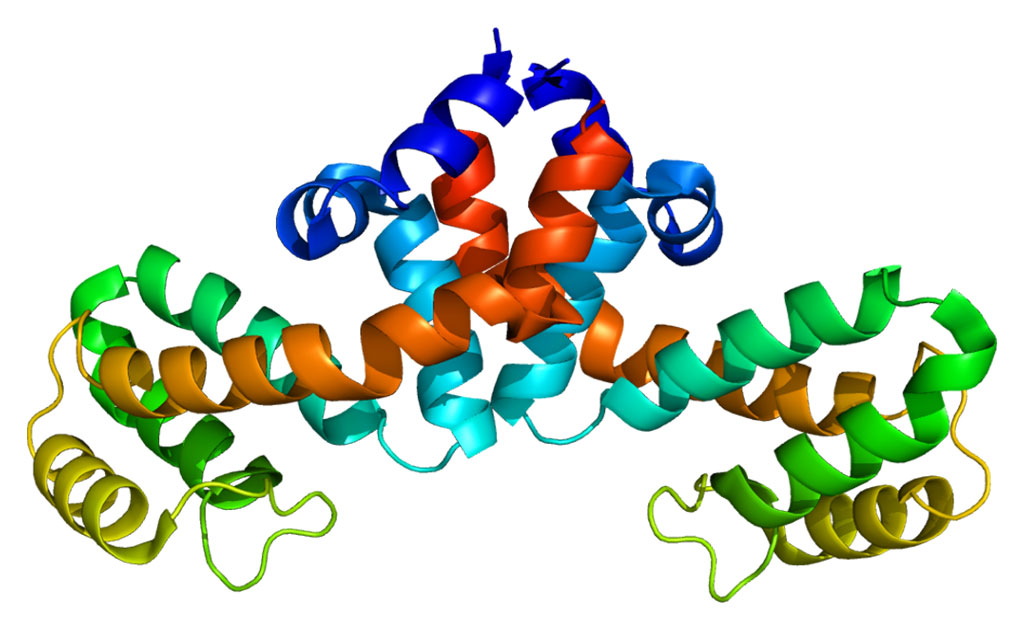
Sensitive X-ray crystallography techniques were used to determine the three-dimensional (3D) molecular structure of the Mycobacterium tuberculosis enzyme RNA polymerase (Mtb RNAP) alone and when bound to the present first-line anti-tuberculosis drug rifampin.Posted on 18 Apr 2017
M. tuberculosis (Mtb) is the causative agent of tuberculosis (TB), which annually kills about 1.8 million people worldwide.
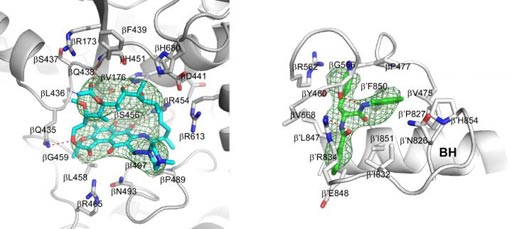
Image: Crystal structures of Mtb RNAP bound to rifampin (left) and Mtb RNAP bound to an AAP (right) show the Mtb RNAP interactions. Gray ribbons, Mtb RNAP backbone; gray, cyan, and green sticks, Mtb RNAP, rifampin, and AAP carbon atoms; red and blue sticks, oxygen and nitrogen atoms; green mesh, mFo-Fc electron density omit map; BH, Mtb RNAP bridge helix (Photo courtesy of Wei Lin and Richard H. Ebright, Rutgers University).
Rifampin is used for the treatment of tuberculosis in combination with other antibiotics, such as pyrazinamide, isoniazid, and ethambutol. For the treatment of tuberculosis, rifampin is administered daily for at least six months. Combination therapy is utilized both to prevent the development of resistance and to shorten the length of treatment. Rifampin inhibits bacterial DNA-dependent RNA synthesis by inhibiting bacterial DNA-dependent RNA polymerase. Resistance to rifampin develops quickly when it is used without another antibiotic. Efforts to circumvent the development of resistance to rifampin have been hampered by the absence of structural information for Mtb RNAP, making rational, structure-based drug discovery for Mtb RNAP impossible.
In the current study, investigators at Rutgers University used X-ray crystallography to establish the crystal structures of Mtb RNAP, alone and in complex with rifampin at 3.8 - 4.4 Angstrom resolution. Results published in April 6, 2017, online edition of the journal Molecular Cell revealed an Mtb-specific structural module of Mtb RNAP and established that rifampin functioned by a steric-occlusion mechanism that prevented extension of RNA.
The investigators also reported the discovery of non-rifampin-related compounds -Nalpha-aroyl-N-aryl-phenylalaninamides (AAPs) - that potently and selectively inhibited Mtb RNAP and Mtb growth, and they described crystal structures of Mtb RNAP in complex with AAPs. AAPs were found to bind to a different site on Mtb RNAP than rifampin, exhibited no cross-resistance with rifampin, functioned additively when co-administered with rifampin, and suppressed resistance emergence when co-administered with rifampin.
"The structure of Mtb RNAP has been the "Holy Grail" for TB drug discovery targeting Mtb RNAP," said senior author Dr. Richard H. Ebright, professor of chemistry and chemical biology at Rutgers University. "AAPs represent an entirely new class of Mtb RNAP inhibitors and are, without question, the most promising Mtb RNAP inhibitors for anti-TB drug development since rifampin. We are very actively pursuing AAPs. We have synthesized and evaluated more than 600 novel AAPs and have identified AAPs with high potencies and favorable intravenous and oral pharmacokinetics."